|
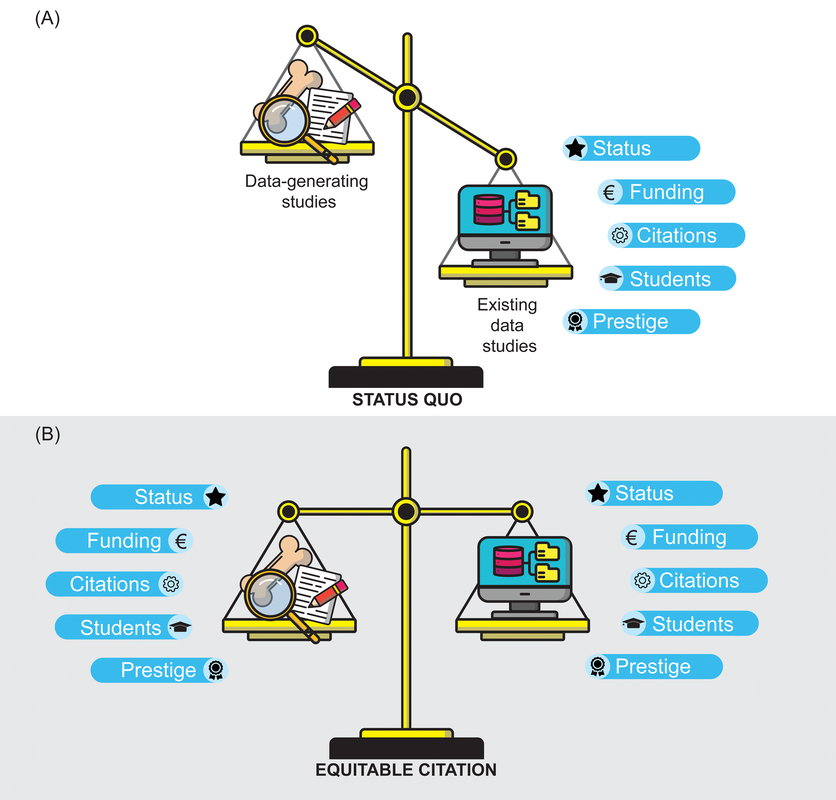
Title: Increasing the equitability of data citation in paleontology: capacity building for the big data future Authors: J.A. Smith, N.B. Raja, T. Clements, D. Dimitrijević, E. M. Dowding, E.M. Dunne, B.M. Gee, P.L. Godoy, E.M. Lombardi, L.P.A. Mulvey, P.S. Nätscher, C.J. Reddin, B. Shirley, R.C.M. Warnock, Á.T. Kocsis Journal: Paleobiology DOI: 10.1017/pab.2023.33 General summary (pulled direct from the non-technical summary available with the paper): Researchers often use large databases to conduct their studies; however, they do not always provide credit, through citations, to the people who produced the data in the databases. In the field of paleontology, researchers use a large database called the Paleobiology Database (PBDB) to study global patterns and processes over millions of years. These studies use data from the PBDB and typically receive a greater number of citations than the original data-producing papers. This creates a situation where the hard work of collecting the data is not credited and rewarded in a fair way, even though this work is equally important to the field of paleontology. By fixing this issue and giving proper credit to data-producing papers, paleontology itself can be strengthened by increasing the incentives for producing data and at the same time creating more high-quality data for everyone to use. Remember your roots Paleontology has historically been a descriptive discipline, focused on describing new fossils, which in turn may represent new species and/or new occurrences (in space and/or time). This is hardly exclusive to paleontology – all natural history disciplines (e.g., geology, other life science sub-disciplines like herpetology or ichthyology) are rooted in simply describing observations. Even today, scientists are constantly reporting new modern occurrences, from new living species (e.g., the Lady Elliott Shrimp Goby) to finding new occurrences, like the deepest known occurrence of a fish below 8,000 m. In paleontology, descriptions of new species and new occurrences are particularly common, even today, because of the incompleteness of the fossil record and the continued erosion and exposure of new fossil-bearing rocks. Such descriptive work is a great example of primary data, which are defined as being unique and novel, or data-generating in the parlance that is used in the above figure (data-provisioning for most of the rest of the paper). This is contrasted with secondary data, which are existing data that are being reused in some form, such as by downloading data from a database that has aggregated primary data from many different sources; a good summary of the distinction can be found here. Because primary data collection involves novel data generation, it is typically (though hardly always) more time-intensive and costly (this should not be misconstrued to mean that secondary data analyses are not or cannot be time-intensive or costly). In paleontology, the official publication of a novel occurrence involves a lot more than just the examination of the fossil and writing it up; it requires someone to have found the fossil to begin with, which is often a resource-intensive process that involves identifying potential fossil-bearing areas, surveying said areas, actually finding a fossil, collecting it, and preparing it. As a result, many data-generating paleontological studies are at a very specific scale (e.g., a single fossil or a set of fossils from one site). Some examples of my primary-data-generating studies from 2023:
The next frontier The vast majority of paleontological work has historically centered on generating primary data because that was the only thing that was really feasible until computers and other technological advancements permitted more comprehensive, and often intensive, analyses that are based on aggregating data together from many sources. Today, such studies are far more tractable and thus far more common, not just for paleontology but across all disciplines (AI being the most prominent recent development that may further expand the research horizons). We also have a variety of openly available databases that aggregate hundreds of thousands of published records, such as the Paleobiology Database (PBDB). A secondary data source could thus be generated in a matter of mere minutes but end up comprising thousands of unique records, each requiring hundreds of hours to have produced. These secondary data studies can take on a variety of forms, such as meta-analyses, bibliometric analyses, or the umbrella term of 'big data' studies; all share a commonality of collating data from many sources in order to create a larger sample size that can be used to tackle questions at larger scales. Some examples from Emma Dunne, one of the other authors on this paper:
A numbers game Academics, like many people, prefer hard numbers (quantitative) to relative assessments (qualitative). Therefore, we often rely on various imperfect numerical metrics and proxies to attempt to assess research (and researcher) quality. Two examples are impact factor (IF) and citation count. Impact factor is a journal-specific metric and is calculated as follows:
Journals with a higher IF are viewed as being more prestigious (selective about what they publish), and even though there are many reasons why a journal's IF does not have much of any bearing on the direct quality of a single article or the researchers who published it, remains a popular metric. For example, Nature and Science, two journals considered among the most prestigious journals across many disciplines, have two-year IFs of 64.8 and 56.9. By comparison, journals like the Journal of Paleontology and the Journal of Vertebrate Paleontology have two-year IFs below 3.0 and are thus considered much less prestigious. Because secondary data studies aggregate data, they can tackle questions that are "bigger picture" and are thus more appealing to selective journals that aren't interested in publishing a description of a new fossil (unless it's a dinosaur or something very cool). As a result, data-generating studies are often published in society journals (e.g., Journal of Paleontology, Acta Palaeontologica Polonica, Journal of Vertebrate Paleontology) that are not considered prestigious. IF is based on citation count, but citation counts can also be used on their own (e.g., a researcher can list the citation counts for all of their articles on their CV) – this can also be used as a semi-quantitative assessment of the quality of research (on the premise that high-quality work is cited more often), but there are also many flaws with correlating citation counts with research(er) quality. The problem
Most researchers are predominantly either data generators or data reusers; this does not mean that one cannot be both, but it is rare for people to be frequently engaged in both forms of analysis such that they are more or less a 50-50 split. Because of the aforementioned reliance on numerical metrics like journal IF and citation counts, researchers with a publishing portfolio that is skewed towards secondary data analysis are more likely to have publications in journals with higher IFs (read as more prestigious) and thus be more competitive / attractive in everything from hiring decisions to grant awards because of a perception that their work is more important, rigorous, "big picture," or applicable when compared to researchers who are primarily data generators publishing in lower-IF journals. However, if primary data generators become increasingly less likely to be funded, the generation of primary data grinds to a halt, and this has downstream effects on what analyses can be done using secondary data when people get stuck with the same old data. Many papers, for example, have demonstrated the growing crisis associated with a lack of funding for taxonomy (e.g., Agnarsson & Kuntner, 2007; Drew, 2011; Löbl et al., 2023). There are a multiplicity of reasons why data generating studies and researchers tend to be undervalued in contemporary academia, but one of them are practices around properly citing sources. As with the citation of previous articles' findings or conclusions, secondary data studies should also be citing the data-generating studies that they rely on for the analysis, yet many fail to properly do so, leading to these gaps in quantitative metrics that lead to the devaluation of data generating research(ers). The data What this study did is put numbers on the qualitative observation that data-generating papers tend to be undercited (undercredited). In paleontology, many of the secondary data analyses rely on aggregating data from the Paleobiology Database, but many of them do not cite the data-generating studies in a way that can be tracked and thus properly credited (I think that some of my own publications have been used in PBDB studies but frankly have no clue which ones). Studies that use the PBDB (secondary data studies) are supposed to "register" in order to receive a PBDB publication number, so these can be easily identified and have their total and per year citation counts obtained/calculated via Google Scholar (orange bar below). From those studies, the data-generating studies that were used by the PBDB studies could be identified (nearly 50,000 unique studies from just 151 PBDB studies for which data could be recovered). The total and per year citation counts for those data-generating studies could then be obtained/calculated via Google Scholar (blue bar below). However, we know that Google Scholar (and every other scholarly aggregator) isn't checking things like Supplemental Information for citaitons, if they exist in the SI to begin with. The number of missed citations (i.e. not tracked by Google Scholar) of data-generating papers could subsequently be calculated from the datasets of those 151 studies (e.g., Smith et al., 2000, was used by 5 PBDB studies) and added to the total citation count (the blue-green bar below). Because not all of the PBDB studies in the focal interval (2001-2021; n=396) had their data recovered, the hidden usage/citation rate of data-generating papers could be extrapolated from the 151 PBDB studies that did have their data recovered, assuming similar usage rates over all 396 studies (the extra blue-green bar below). These extrapolated numbers result in a mean citation rate that is nearly the same as that of PBDB studies, rather than the presently recorded rate that is only about one-third of the rate. 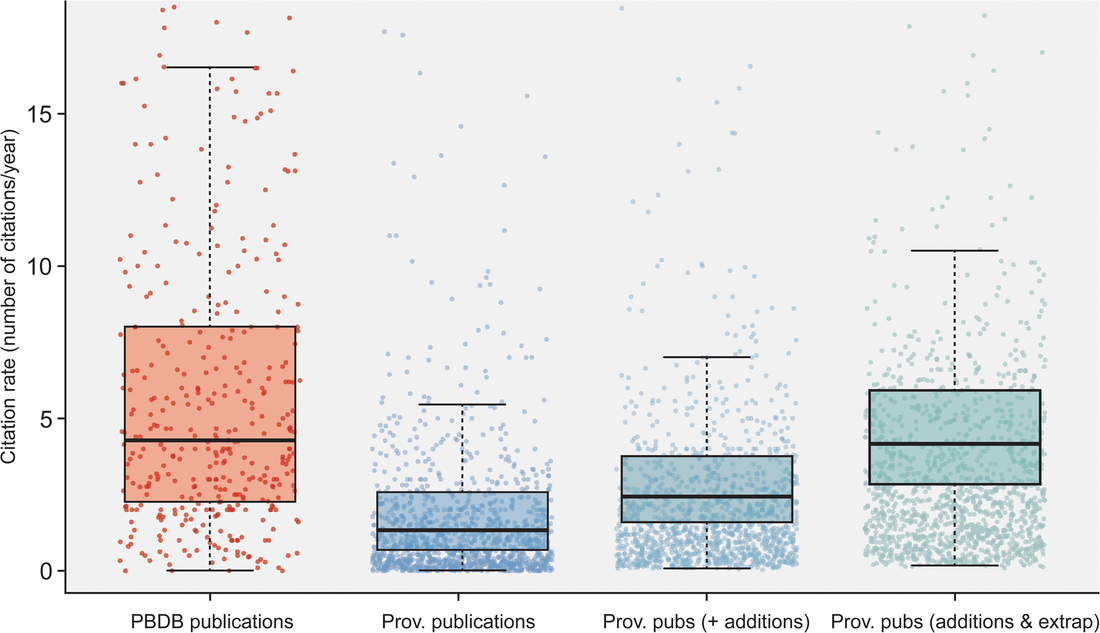 Figure 2 from the paper: "Citation rates for official Paleobiology Database (PBDB) publications and the data-provisioning publications used in those PBDB publications. Only data-provisioning publications from the same time frame (since 2001) as PBDB publications are included to standardize for temporal effects. Citations to data-provisioning publications (i.e., primary literature) are presented as the current rate (i.e., no additions for neglected citations), the projected rate when including citations from PBDB publications where data were available (k = 112; i.e., additions), and the projected rate when making those additions and extrapolating to the entire set of PBDB publications (k = 396; i.e., additions and extrapolated)." Predictably, if the true citation count of data-generating papers can be demonstrated to be higher than what is presently tracked across scholarly aggregators like Google Scholar, the journals in which these papers are typically published should also have higher impact factors than what is presently reported, and this is shown below (see the discussion in the paper for more context about different year-intervals in impact factor calculations and other historical conditions for publishing). Collectively, these analyses provide clear evidence that data-generating studies are systemically undercited, which can produce misleading perceptions of interest in and quality of these studies and the journals they are published in (even if that is in large part because we rely too much on flawed metrics to assess these). 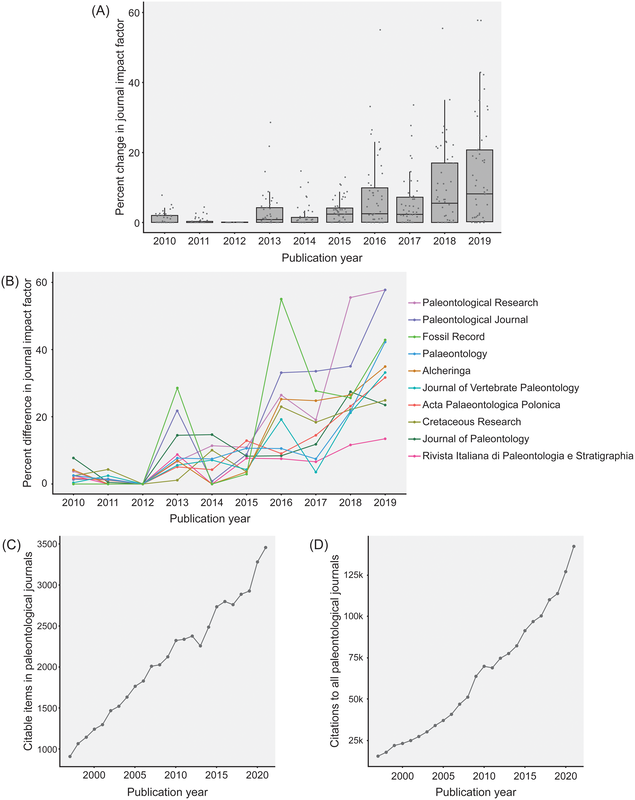 Figure 3 from the paper: "The effects of adding neglected citations from data reuse on journal impact factor (JIF; A, B) and general patterns in publishing trends in paleontology (C, D). A, The increase in JIF for the 55 journals categorized to paleontology by Clarivate, for the period of 2010 to 2019. Note, an outlier value of 172% in 2018 for PalZ was not plotted. B, Increases in JIF for the 10 paleontological journals most affected by neglected citations, only including those with complete data for the duration of 2010 to 2019. For raw data for all 55 paleontological journals from 1997 to 2021, see “7_paleo_journal_JIFcalculation.csv” in Smith et al. (2023a). C, The number of citable items published in paleontological journals each year. D, The number of citations to items published in paleontological journals each year." Don't hate the player, hate the game A lack of proper citation of data-generating studies by data-reusing studies can and certainly does result from poor research practices around citation (I often see people remove license/copyright and citation information from database downloads in my day job), but even well-intentioned researchers can frequently have their hands tied by various other factors, mainly on the end of the journals that we publish in. For example, these are some policies found across various journals that can lead to a lack of citations of primary data:
A way forward Many of these hindrances are structural at the level of a journal or publisher, so it may seem that there is little to be done to more accurately and equitably track data reuse. However, there are a variety of actions that authors can take now to try and work around these existing limitations. One example:
Finally, the biggest challenges lie in advocating for structural changes. For example, as research output rises, the incentives to perform peer review, which is practically never compensated or rewarded in any form, decline. Peer review is the primary mechanism that should be responsible for ensuring rigor of a study, which includes proper data sharing and data citation, but there is not much incentive for reviewers to even check supplemental information files if doing so would require substantial time investment. Other target areas include advocating for changes in journal policies in how citations can be provided in a format that is discoverable and indexed by the aggregators that we all rely on to track citations is the most crucial and alternative means of evaluating research quality and performance. A few journals (e.g., Global Ecology & Biogeography) do allow for references cited only in Supplemental Information to be listed in the main-text References so that they are picked up by indexers. As journal editorial boards are made up of normal (well, somewhat normal) researchers, they also have some ability to direct journal policies in a variety of ways, from formatting prescriptions to journal scope. Development and adoption of other metrics of tracking and recognizing scholarly contributions in merit-based processes, in which "normal" researchers are also directly involved, will also be critical.
0 Comments
Title: Triassic stem caecilian supports dissorophoid origin of living amphibians Authors: B.T. Kligman, B.M. Gee, A.M. Marsh, S.J. Nesbitt, M.E. Smith, W.G. Parker, & M.R. Stocker Journal: Nature DOI: 10.1038/s41586-022-05646-5 General summary: The origin of modern amphibians (frogs/toads, salamanders/newts, caecilians), which are more often termed 'lissamphibians' by scientists to differentiate from the more ambiguous 'amphibians,' has long been a vexing problem. The three modern groups are remarkably morphologically disparate, which makes it hard to both confidently identify and conceive of the ancestral lissamphibian. Lissamphibians today are also relatively small, a pattern thought to have characterized much of their evolutionary history and also an attribute that predisposes their remains to not have been fossilized. Exacerbating this is the extremely poor record of caecilians, which, even today, are a rather cryptic group found only in the equatorial regions and that typically burrow, making them rather hard to observe. Burrowing animals, especially those that not only burrow but that spend much of their lives underground, also have a poor fossil record, which compounds the problems for caecilians. The first caecilian fossils were not even reported until 1972 (previous reports were a misidentifed catfish spine and a misidentified cephalopid, respectively), and to date, there are less than a dozen distinct occurrences of fossil caecilians known globally over an 180 million year interval.
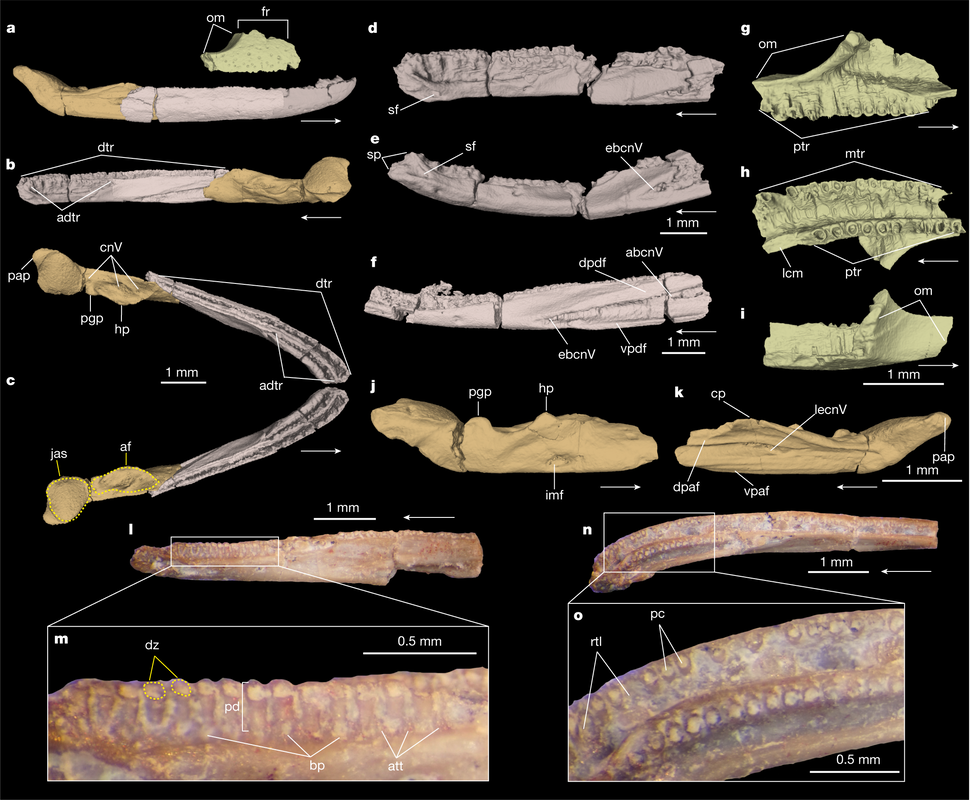
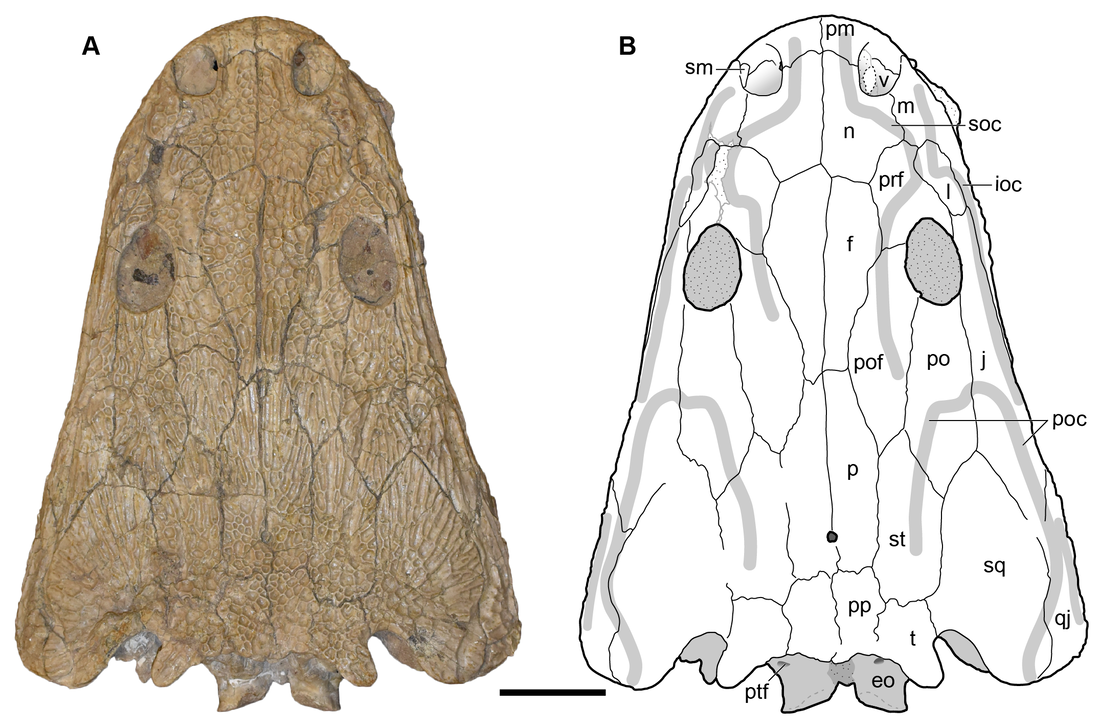
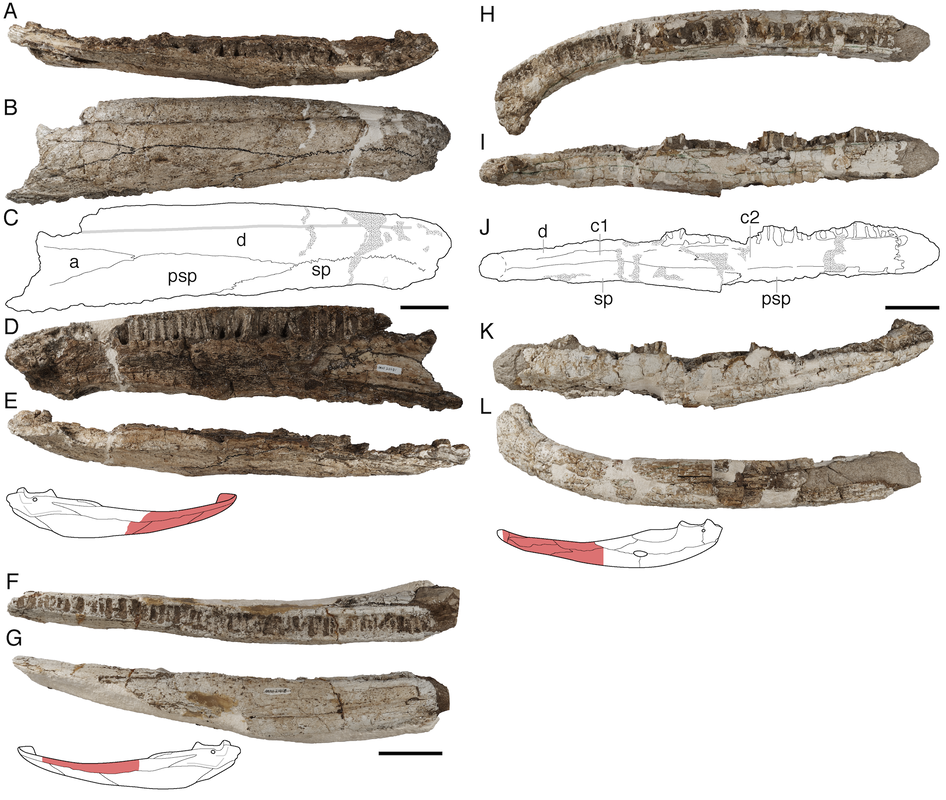
In this study, led by my colleague and current Virginia Tech PhD student Ben Kligman, we report nearly 100 new specimens of the earliest known caecilian in the fossil record from a single Late Triassic site in Petrified Forest National Park, Arizona. Although no complete skulls or skeletons are known, numerous fragments preserve unequivocally diagnostic features found only in caecilians, such as a jaw comprised of largely fused elements that remain separate in other tetrapods and two rows of small teeth with a distinctive feature called pedicelly – a dividing zone at the mid-height of the tooth that often leads the tips to be lost during preservation. This occurrences pre-dates the previous oldest occurrence of caecilians by at least 35 million years and provides new insights into the early stages of the group's evolution. In particular, the new fossils appear to capture the transition toward the modern caecilian condition in which there is extensive co-ossification of multiple elements to form a more consolidated skull (good for burrowing). Their occurrence in Arizona, which was positioned close to the equator in the Late Triassic, suggests that an origin within the equatorial belt also constrained their dispersal, therein offering an explanation as to why caecilians remain tied to these regions when frogs and salamanders have nearly a global distribution except at the poles (and Australia for salamanders). Title: Revision of the Late Triassic metoposaurid “Metoposaurus” bakeri (Amphibia: Temnospondyli) from Texas, USA and a phylogenetic analysis of the Metoposauridae Authors: B.M. Gee; A.M. Kufner Journal: PeerJ DOI: 10.7717/peerj.14065 General summary: Frequent readers of this blog are of course familiar with my love for metoposaurids, one of the most iconic groups of North American temnospondyls. It is no secret that metoposaurids were my "gateway drug" to the world of terrifyingly large and unrecognizable amphibians, and so they always have a special place in my heart. My previous research has largely focused on two of the three species, Anaschisma browni (ex. Koskinonodon perfectus, ex. Buettneria perfecta) and Apachesaurus gregorii, which are the two most common metoposaurids that we get in the Late Triassic of North America. If you go to a museum and see a metoposaurid (most major museums in the U.S. have a metoposaurid on display), it's likely Anaschisma browni, although the label may be two or three junior synonyms out of date. The third species, known only from two sites in Texas and one site in Nova Scotia, doesn't get as much press - it was last (re)described in 1932! Ninety years later, my buddy Aaron Kufner and I are pleased to produce a full redescription of this taxon, long referred to as 'Metoposaurus' bakeri, based on a reexamination of material from the type locality that now lives at the University of Michigan Museum of Paleontology (UMMP) collections in Ann Arbor. With a thorough redescription spanning 134 print pages (probably too thorough, even by my own standards), we provide extensive documentation of the skeletal anatomy of this species (1930s-era drawings have their limitations), conducted more phylogenetics analyses to test the relationships of metoposaurids (this has gotten no better since the 1930s), and ultimately concluded that this species cannot be placed in an existing genus like Metoposaurus (otherwise only known from Europe). To that end, we created a new genus name, the mouthful Buettnererpeton, which honors a longtime fossil preparator at the UMMP, William H. Buettner, who worked extensively with E.C. Case, the museum curator who named the species in 1931. The suffix comes from -herpeton, meaning 'creeping animal' in Greek, which is a common component of names of early reptiles and amphibians. Our taxonomic act has implications for biostratigraphy (relating distantly situated rocks based on which taxa occur in them), both globally and within North America, and we discuss everything from future work needed on metoposaurids to why their phylogenetic relationships are so badly resolved. This is a 'boundary-crossing' project that originated when I was a Ph.D. student in the summer of 2019 and has only now made it to the finish line, so it is particularly memorable for me in that regard.
Title: Cold capitosaurs and polar plagiosaurs: new temnospondyl records from the upper Fremouw Formation (Middle Triassic) of Antarctica Authors: B.M. Gee; C.A. Sidor Journal: Journal of Vertebrate Paleontology DOI: 10.1080/02724634.2021.1998086 General summary: The Middle Triassic captures a diverse global record of temnospondyls, which is also when we start to see the pinnacle of the evolution of large body size, with many taxa routinely exceeding skull lengths of half a meter and body lengths of probably 2m or greater. A variety of different groups are present at this time, all of which appeared in the Early Triassic and which would also continue through the Late Triassic, and many ecosystems were host to several different species of no close relatedness. However, the Antarctic record of Middle Triassic temnospondyls has only comprised members of a single clade, the capitosaurs. Dated and brief historical notes suggested the possible presence of another clade, the long-snouted crocodilian-like trematosaurs, but this was never substantiated, and thus the Antarctic record, despite preserving at least three different species, captures an overall much lower diversity of temnospondyls than found elsewhere around the world. In this study, we took a look at some of this more ambiguous historical material, combined with more recently collected material of some very large lower jaws. While every single one of these lower jaws belongs to a capitosaur, there is a partial interclavicle (part of the shoulder girdle) that is very clearly not that of a capitosaur but instead that of a plagiosaurid, a peculiar short-snouted clade that has hundreds of records from the northern hemisphere but a mere two others from the southern hemisphere (both of those are from the Early Triassic). We speculate on some of the reasons why the Antarctic record, which is undoubtedly undersampled, might reflect real patterns of differing ecologies among large-bodied temnospondyls (i.e. 'big crocodile analogue' is a gross oversimplication).
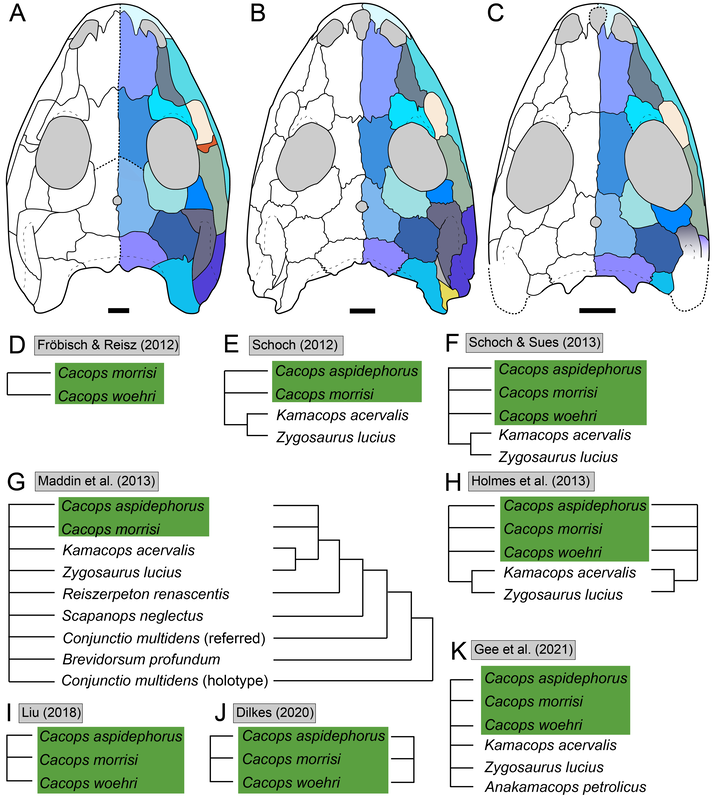
Title: Returning to the roots: resolution, reproducibility, and robusticity in the phylogenetic inference of Dissorophidae (Amphibia: Temnospondyli) Authors: B.M. Gee Journal: PeerJ DOI: 10.7717/peerj.12423 General summary: Phylogenetics, the means of inferring relationships among organisms and one way of classifying them, is a timeless part of paleontology, in part because our datasets are much more limited than those for modern organisms in which genetic barcoding is both widely accessible and robustly informative. As a result, there is often a lot more uncertainty and disparity among different analyses, and people should generally be skeptical when someone says there is a strong consensus for a given group, no matter how well-known (paleo nerds will definitely recall the debate a few years back about what 'Dinosauria' constituted). Temnospondyls are generally not well-known, and this means that they are not as well-studied. This often has the effect of resulting in one or two people producing a lot of the work for a given group, which can lead to an overstated consensus when the same dataset keeps getting used. Dissorophids, the armoured dissorophoids, are a great example of this. There is a ton of work, but pretty much everyone uses the same phylogenetic matrix. As you might imagine, analyzing pretty much the same dataset a bunch of times should lead to pretty much the same result, which would not be particularly compelling as a "consensus," but, SPOILER, there is in fact no consensus among these nearly identical analyses! That's a huge issue. This study delves into why, both through a separate phylogenetic matrix and from closely scrutinizing the previous one. There is a ton of data packed into it, but the short and skinny are that (1) previous studies have been compromised by systemic errors from the original; and (2) there is a general lack of resolution / of well-supported nodes among dissorophids (i.e. we don't know a lot more than we do know). Then there's a bunch of other toss-ins like missing holotypes and chimeras. Keep reading to find out more!
|
About the blogA blog on all things temnospondyl written by someone who spends too much time thinking about them. Covers all aspects of temnospondyl paleobiology and ongoing research (not just mine). Categories
All
Archives
January 2024
|
 RSS Feed
RSS Feed
